Data Quality Report configuration
This document was built from version 0.1.14, Sep 25, 2023 at 22:36
Introduction
The current versions of this document are available online as HTML and PDF. These are built automatically when new versions are merged.
Type |
Link |
|---|---|
HTML user’s manual |
https://o4-dqr.docs.ligo.org/o4-dqr-configuration/index.html |
PDF user’s manual |
https://o4-dqr.docs.ligo.org/o4-dqr-configuration/dqrconfiguration.pdf |
HTML programming docs |
https://o4-dqr.docs.ligo.org/o4-dqr-configuration/api/index.html |
PDF API |
https://o4-dqr.docs.ligo.org/o4-dqr-configuration/api/taskmanagerapi.pdf |
This project has a few applications that manage DQR configuration files. The configuration processing is designed to make it easy to add new analyses to the report, defining all requirements from the infrastructure in the configuration file.
Current status
Version 0.1.14 is the current version it will create htCondor DAG and submit files from configuration files. It should be considered a prototype whose purpose is demonstrate feasibility and evoke suggestions.
Download
This software is available as a pip installable, pure python package. It is available from git.ligo.org with appropriate LVK credentials. Version 1.0.x will be open source and moved to github when reviewed and ready for O4. Use the command:
git clone git@git.ligo.org:o4-dqr/o4-dqr-configuration.git
or
git clone https://git.ligo.org/o4-dqr/o4-dqr-configuration.git
Contributions
We welcome any and all help in this project. We ask that all contributions be made in a branch or fork then submitted with a Merge Request – see How to contribute to the TaskManager project for details.
Installation
The prototype project has not been uploaded to PyPI and Conda so please clone the git project (see above). It is designed to be installed into a recent IGWN conda environment see https://computing.docs.ligo.org/conda/
We recommend using a development conda environment be used to isolate any version conflicts from production
conda activate igwn-py38
cd o4-dqr-configuration
pip install .
We can also work with a minimal Conda environment using Python 3.7 or 3.8. To create a new environment a conda or miniconda installation is needed. If neither is installed mini conda is recommended see https://conda.io/projects/conda/en/latest/user-guide/install/index.html
To install a minimal environment:
conda create --name o4_dqr_proto python=3.8
conda activate o4_dqr_proto
conda install m2crypto pykerberos
# For sphinx documentation building
conda install sphinx sphinx_rtd_theme
pip install git+ssh://git@git.ligo.org/o4-dqr/o4-dqr-configuration.git
Building the documentation
cd docs
make html
The main html page will be build/html/index.html.
If you have texlive installed you can create a pdf version with
cd docs
make latexpdf
The single PDF will be build/latex/dqrconfiguration.pdf.
Application: dqr-create-dag - processing events step 1
The dqr-create-dag application uses configurations and command line options to create htCondor DAG files and a directory structure for processing events in GraceDB. We can also run some tasks at specific GPS times. This depends on whether the task is designed to not rely on information unique to GraceDB.
$ dqr-create-dag --help
usage: dqr_create_dag [-h] [-v] [-V] [-q] [--ev-json [EV_JSON ...]]
[-g [GRACEID ...]] [--ev-file [EV_FILE ...]]
[--submit-dag] -c CONFIG_FILES [CONFIG_FILES ...]
[--config-overrides [CONFIG_OVERRIDES ...]]
[--config-dir CONFIG_DIR] [--max-far MAX_FAR]
[--no-globus]
Create a Condor DAG to process one or more GraceDB [super]events
optional arguments:
-h, --help show this help message and exit
-v, --verbose increase verbose output
-V, --version show program's version number and exit
-q, --quiet show only fatal errors
--ev-json [EV_JSON ...]
One or more json files with gracedb inormation
-g [GRACEID ...], --graceid [GRACEID ...]
One or more GraceIDs or GPS times.
--ev-file [EV_FILE ...]
File with one event specifier per line. It maybe a
GraceId, GPS time, or path to file with .json
extension
--submit-dag submit the dag on this machine
Configuration:
-c CONFIG_FILES [CONFIG_FILES ...], --config-files CONFIG_FILES [CONFIG_FILES ...]
One or more configuration files, loaded in order
--config-overrides [CONFIG_OVERRIDES ...]
override one or more configuration options. Format is
<section>[:<key>[=<value>]] If no value is specified
key is removed from specified section. If no key is
specified section is removed.
--config-dir CONFIG_DIR
Default directory for configurationfiles. This only
affects relative pathson the command line or include
directives.
--max-far MAX_FAR Maximum false alarm rate to process
--no-globus
Event Identification
There are 3 ways to identify an event to process:
GraceID is the event or superevent name in GraceDB. Some of these are public which do not need authorization to access. Private events need login credentials.
GPS time, a floating point number definining the time to analyze. Note some tasks may require more information and may reject these requests.
A JSON file created from a GraceDB
GraceID and GPS times are listed on the command line with the –graceid (-g) option. The json files are specified on the command line as a path with the –ev-json option. The path may be absolute or relative to the current working directory.
The –ev-file option allows a list of events so be listed in a file. The option allows easy testing of the analyses on a large number of events. The format of the file is
One event per line.
A comment starts with a hash (#). It may be on a line by itself or following an event.
- An event can be specified by
A GraceID: a string that doesn’t look like a floating point number and does not end with “.json”
A GPS time: a floating point number.
A json file specifier: a path to an existing readable file. The path must end with a “.json” extension. It may be absolute, starting with a slash (/). If it’s a relative path, the current working directory is checked first, then the directory with with the event file.
Blank lines are ignored.
Application gdb2json - Copy GraceDB event to a file
The gdb2json application is used to copy information so that processing can be done without access to GraceDB, such as during the Continuous Integration testing we do with gitlab.
$ gdb2json --help
usage: gdb2json [-h] [-v] [-V] [-q] [-o OUTDIR] [-i INFILE] [graceid ...]
Copy GraqceDB info to JSON fie
positional arguments:
graceid One or more graceids
optional arguments:
-h, --help show this help message and exit
-v, --verbose increase verbose output
-V, --version show program's version number and exit
-q, --quiet show only fatal errors
-o OUTDIR, --outdir OUTDIR
Path to directory for output JSON files
-i INFILE, --infile INFILE
File with list of graceids
Configuration syntax
Configuration file basics
The configuration for DQR is very flexible, designed to make task development as easy as possible.
An example configuration file:
[general]
# some system descriptors
output_dir = ${HOME}/public_html/events
output_url = http://${HOSTNAME}/~${USER}/events/
gracedb_url = https://gracedb.ligo.org/api/
deep_variables = mchirp ttotal template_duration sigmasq
[auth]
# Specify a robot keytab if needed to access GraceDB and/or dqsegdb
keyname = dqr/robot/dqr.ligo.caltech.edu@LIGO.ORG
keytab = ${HOME}/.private/dqr_robot_dqr.ligo.caltech.edu.keytab
min_cert_time = 2
cert_directory = ${HOME}/.private
[condor]
# default classads for condor submit files
accounting_group = ligo.${run_type}.o3.detchar.transient.dqr
accounting_group_user = joseph.areeda
getenv = True
request_memory = 500MB
request_cpus = 1
Some basics:
The default extension for config files is .ini
Files may contain multiple sections
Keys are case sensitive
Keys are unique within a section. If there are duplicates, the last value is used
If values contain spaces, they should be quoted.
For DQR section names should follow identifier syntax
[_:Alpha][_:Alphanum]* (Must start with a letter or underscore (_) followed by letters, numbers or underscore
Section names are used as Job identifiers in the Condor DAG
The command line for applications that use the DQRConfig class must specify at least one configuration file.
The –config-dir command line parameter specifies a root path for any relative path in the command line or in include and include_dir directives.
The –config-overrides command line parameter provides a convenient way to adjust a configuration parameter or two. For example, given the config file above, suppose we want to test with different condor defaults. We could add the following to the command line:
--config-overrides "DEFAULT:include = ${HOME}/test/condor_new.ini"
Special sections and special variables
There are some special sections but most define variables that are used in creating commands for the analyses. There are multiple levels of scope
Global, used in all levels
Per event, includes any information from GraceDB
Condor defaults
Iterator definitions
Authorization, robot keytab information [optional]
Specific tasks
Special sections
[DEFAULT]
Defined by the section name, the Python ConfigParser treats every member of this section as a member of every other sections, including the executable sections used to create submit files. Be careful with items in this section: they can invalidate a submit file.
[condor]
Defined by the section name these members are added to submit files unless overridden.
executable
Defined by a member “executable”. These sections represent the individual analysis tasks and are used to create the Condor submit files.
iterator
Defined by having a member named “iterator”. These sections are used with the “iterate” directive in an executable section. See the Iterators section below.
auth
Identifies the path to a robot keytab, and the principal name that can be used to create a Kerberos TGT. The robot TGT can be used to create an x509 certificate. See the Authorization section below.
Note
If this section is not available or the command line option --noglobus is used no
ticket or certificate will be created. Thus if access to authenticated services
such as NDS2, GraceDB or dqsegdb are needed it is the task’s responsibility.
General section variables
output_dir
A path to the directory where results are saved. This is usually a directory available to a webserver such as ~/public_html/dqr This field often uses one of the time variables see (:ref: date_strings_for_subdirectory_names) to organize results by the date of the event or the date it was processed. For each event a directory is created and inside that directory is a subdirectory for each analysis run. In the top level directory is a symbolic link to the latest analysis. For example:
~/public_html/dqr-events ├── [ 22] S123456 -> S123456_dir/S123456_02 └── [ 128] S123456_dir ├── [ 64] S123456_01 └── [ 64] S123456_02
output_url
A URL that points to the output directory specified for example: https://ldas-jobs.ligo.caltech.edu/~dqr/dqr-events The appropriate link will be generated from this base to annotate the GraceDB superevent.
gracedb_url
URL used to access GraceDB to get information about an event or to update the event with information about our results, usually a web link to our report. It defaults to: https://gracedb.ligo.org/api/
keyname and keytab
information needed to get a Kerberos TGT and x509, (sci-token?) with a robot keytab
deep_variables
A deep variable is one that is not at the top level of a GraceDB event structure. For example gstllal events may have an array of single inspiral parameters, one per interferometer. The parameters of the matching template are stored here.
The default list of deep variables include: eff_distance, mass1, mtotal, spin1z, template_duration, mchirp
Since these variables are not in fixed locations it is difficult to specify them exactly. The “extra variable” section of the GraceDB event will be searched for each variable specified. For example:
deep_variables = mchirp gamma2 f_final
Note
If there are multiple instances of a specified variable the first value found will be used. This may override a top level variable.
describers
A space separated list of variable names in any section that affects keys in the executable task definitions. If one of the variable names is used as a key in the condor section or an executable section, the line will be copied to the submit file verbatim, preceded with a plus (+).
The default describers list is “description librarian question” describers in the configuration files add to the list.
For example:
[condor]
describers = AccountingGroup Experiment
getenv = True
[status]
description = get status around the trigger time
librarian = Virgo DetChar group (detchar@ego-gw.it)
Experiment = detchar
executable = get_status
arguments = ${t_0}
Would produce a submit file like:
arguments = 1253922430.9494
error = /Users/areeda/public_html/events/1909/S190930ak_16/omega_job2/condor-omega-S190930ak.err
executable = get_status
getenv = True
log = /Users/areeda/public_html/events/1909/S190930ak_16/omega_job2/condor-omega-S190930ak.log
output = /Users/areeda/public_html/events/1909/S190930ak_16/omega_job2/condor-omega-S190930ak.out
+description = get status around the trigger time
+librarian = Virgo DetChar group (detchar@ego-gw.it)
+Experiment = detchar
queue 1
tier_1_universe
Tier 1 job are low latency and if the task manager is run on dedicated resources the “local” universe has the least delay. The option is any valid htCondor universe such as vanilla
condor_extra
In a task definition section this parameter is a multi-line string that is added verbatim to the submit file
Conditional statements
There are a few conditional tests that are expressed as a variable assignment. These are used in an executable section to determine if that task will be included in the DAG
Date strings for subdirectory names
These strings are based on the event name: for example, S200305ay would produce “2003”. If the event is defined by GPS time and no Grace ID is available, the GPS time is used. There are also strings available based on when the program is run.
These can be used to organize results by the event date or the date the report is run. For example:
output_dir = ${HOME}/public_html/events/${ev_yymm}
output_url = http://${HOSTNAME}:~${USER}/events/${ev_yymm}
ev_yymm |
Derived from grace ID if available, otherwise from t_0 |
ev_yyyymm |
Derived from t_0 |
now_yymm |
Derived from current time (UTC) |
now_yyyymm |
Derived from current time (UTC) |
Special section for Condor submit files
The condor section contains the default classads for every submit files. Each task may override these or add to them. An example:
[condor]
accounting_group = ligo.${run_type}.o3.detchar.transient.dqr
accounting_group_user = joseph.areeda
getenv = True
request_memory = 500MB
request_cpus = 1
Iterators
There are instances where a specific job is repeated based on a variable such as interferometer. An example would be to create an Omega scan for each interferometer at the event t0
To implement this, a special section is defined as having a key named iterator defined, the value of which is ignored. In a task definition a key named iterate with the value being a list of iterator section name. For example:
[v1]
iterator = 1
ifo = V1
frame_type = V_HrecOnline
channel = V1:h_16384Hz
[l1]
iterator = 1
ifo = L1
frame_type = L-L1_llhoft
channel = L1:GDS-CALIB_STRAIN_CLEAN
[t0_omega]
iterate = l1 v1
executable = ${python_igwn38}/gwpy-plot
arguments = gwpy-plot qtransform --chan ${channel} --gps ${t_0} --out ${outdir}/t0_omega_${ifo}.png
This will produce two submit files, and two output directories, one for each ifo. In the above example the directories t0_omega_l1 and to_omega_v1 hold the submit files and the ${outdir} variable points to the respective subdirectory.
Dependent tasks
There are situations where one task needs access to the output of another (parent) task. Using the parent key in an executable section ensures that the parent will complete successfully before the child is started. See HTCondor DAGman.
The create DAG operations offer two ways of finding the directory of the parent task.
Explicitly define the subdirectory for one or more tasks.
Deduce the output directory path.
The directory structure that holds evertyhing for the analysis of an event is shown in the nearby figure.
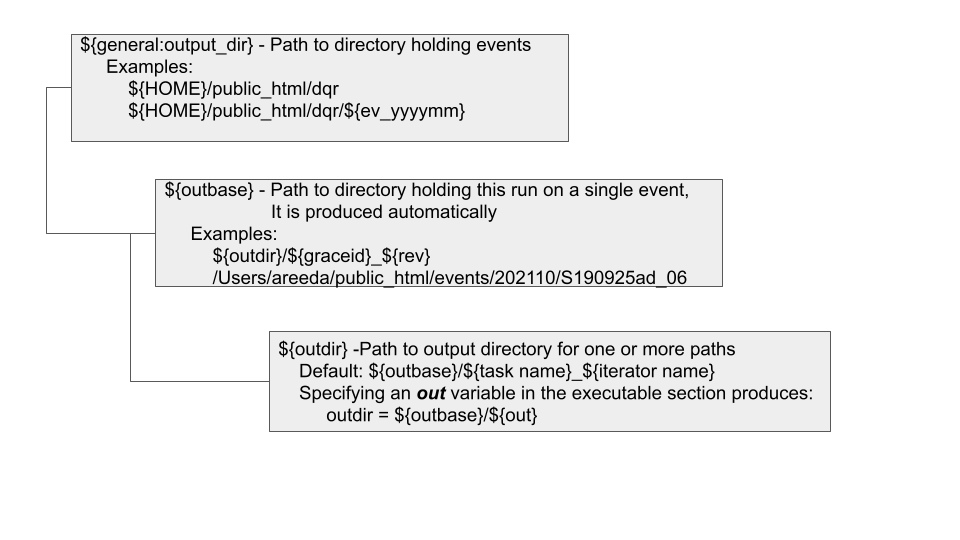

Fig. 1 DQR directory structure
In the configuration file, the outdir variable must be defined. This defines the top level directory for this run. Beneath that directory are the subdirectories for each event. If an event is processed more than once under the same outdir, a revision number is appended. An internal variable named outbase is created which points the directory holding this DAG and all tasks results, log files, submit files.
Note
If you need to access to the results of a task, make it a parent of your task to ensure it completeness before your task runs.
The configuration below is an example of parallel tasks used to create Omega plots, followed by a script to combine the resulting images into a montage.
The t0_omega section uses the iterator key to produce 2 tasks which run in parallel in different htCondor slots. These jobs are named t0_omega_h1 and t0_omega_l1 from the <section_name>_<iterator> pattern
The out key specifies the output directory ${outdir} as t0_omega_dir. This allows the different tasks to specify the same directory on their command line.
The parent key identifies the job (irrespective of the directory) including the iterators:
parent = t0_omega_l1 t0_omega_h1
How to specify directory of a task
It is often useful to use the output of an unrelated task as input to a new task. The directory of the parent task is always a subdirectory of the ${outbase} path variable. The subdirectory name then can be determine by one of these rules:
If the task’s section contains an out key, the subdirectory will be ${outbase}/<value of out key>
If the task does not have an iterate key, then the the subdirectory will be ${outbase}/<task name>
If the task has an iterate key, there is a subdirectory for each iterator identfied by ${outbase}/<task name>_<iterator>
Sample parallel task configuration:
# test looping over variable sets (iterators)
[v1]
iterator = 1
ifo = V1
frame_type = V_HrecOnline
channel = V1:h_16384Hz
[l1]
iterator = 1
ifo = L1
frame_type = L-L1_llhoft
channel = L1:GDS-CALIB_STRAIN_CLEAN
[h1]
iterator = 1
ifo = H1
frame_type = H-H1_llhoft
channel = H1:GDS-CALIB_STRAIN_CLEAN
[t0_omega]
iterate = l1 h1
tier = 1
out = t0_omega_dir
executable = ${igwn_bin}/gwpy-plot
arguments = qtransform --chan ${channel} --gps ${t_0} --out ${outdir}/
request_memory = 1500M
[omega_montage]
tier = 1
description = merge existing images
librarian = joseph.areeda@ligo.org
include_in_dag = True
out = t0_omega_dir
executable = ${dqr_bin}/mkmontage.sh
parent = t0_omega_l1 t0_omega_h1
Common issues with htCondor solutions
Anticipated failure in parent job
Consider the iterator example above. We can define iterators for each interferometer with the understanding that we must run the child task even if one or more parent tasks fail. The default Dagman behavior is to cancel a child task if any parent task indicates failure with a non-zero return code. A simple way to deal with this is to use a POST script in the t0_omega section: we can add the following line to ignore the return code.
SCRIPT POST t0_omega /usr/local/bin/bash -c exit 0
Task held because requested memory exceeded
Estimating how much memory a job will need is not an easy task. Choosing an upper limit is not unreasonable, but requesting unneeded resources impacts the efficiency of the whole cluster.
An easy way to measure memory usage is to use the command
/usr/bin/time -v <command>
It is slighly different on MacOS
/usr/bin/time -l <command>
Note
/usr/bin/time is not the same as the built-in bash command time.
Another issue is that memory requirements are often data-dependent with some runs requiring significantly more. A good way to deal with data dependencies and unclear limits is to start with a reasonable request, then to respond automatically by increasing memory limits and releasing the hold. This can be accomlished wite the following classads:
request_memory = ifthenelse(isUndefined(MemoryUsage),2000,3*MemoryUsage)
periodic_release = (HoldReason == 26) && (JobStatus == 5)
This works by setting the initial memory request to 2 GB, the if the job is held (JobStatus == 5) and the reason is because memory request is exceeded (HoldReason == 26) then the requested memory is trippled and the hold released.
Other variables
In addition to variables defined in the configuration environment variables and those defined in a GraceDB [super]event may be used.
Note the environment variables are examined while the DAG is being created.
The GraceDB variables are dependent on which pipeline submitted the event.
Example task configuration
[overflow_check]
description = uses dqsegdb to plot overlows detected
librarian = adrian.helmling-cornell@ligo.org
include_in_dag = True
tier = 1
question = Are known sources of noise without auxiliary witnesses active?
parent = segments
executable = ${python_igwn38}
request_memory = 400MB
arguments = "-m overflow.overflow_check -vvv --out ${outdir} ${graceid}"
This produces a submit file named something like condor-overflow_check-S190930ak.submit:
$cat condor-overflow_check-S190930ak.submit
+description = "uses dqsegdb to plot overlows detected"
+librarian = "adrian.helmling-cornell@ligo.org"
+question = "Are known sources of noise without auxiliary witnesses active?"
accounting_group = ligo.dev.o3.detchar.transient.dqr
accounting_group_user = joseph.areeda
arguments = "-m overflow.overflow_check -vvv --out /home/areeda/public_html/events/S190930ak/overflow_check S190930ak"
error = /home/areeda/public_html/events/S190930ak/overflow_check/condor-overflow_check-S190930ak.err
executable = /home/areeda/miniconda3/envs/igwn-py38/bin/python
getenv = True
log = /home/areeda/public_html/events/S190930ak/overflow_check/condor-overflow_check-S190930ak.log
output = /home/areeda/public_html/events/S190930ak/overflow_check/condor-overflow_check-S190930ak.out
request_cpus = 1
request_memory = 400MB
universe = local
queue 1
How to contribute to the TaskManager project
The TaskManager project uses a standard fork-branch-review-merge workflow. This is a [very] brief overview of the git operations you may want to use.
The repository for the project is located at https://git.ligo.org/o4-dqr/o4-dqr-configuration
If you plan to contribute to the code, please fork the project by using the fork button in the upper right. If you do not plan to contribute to the code, feel free to clone it instead.
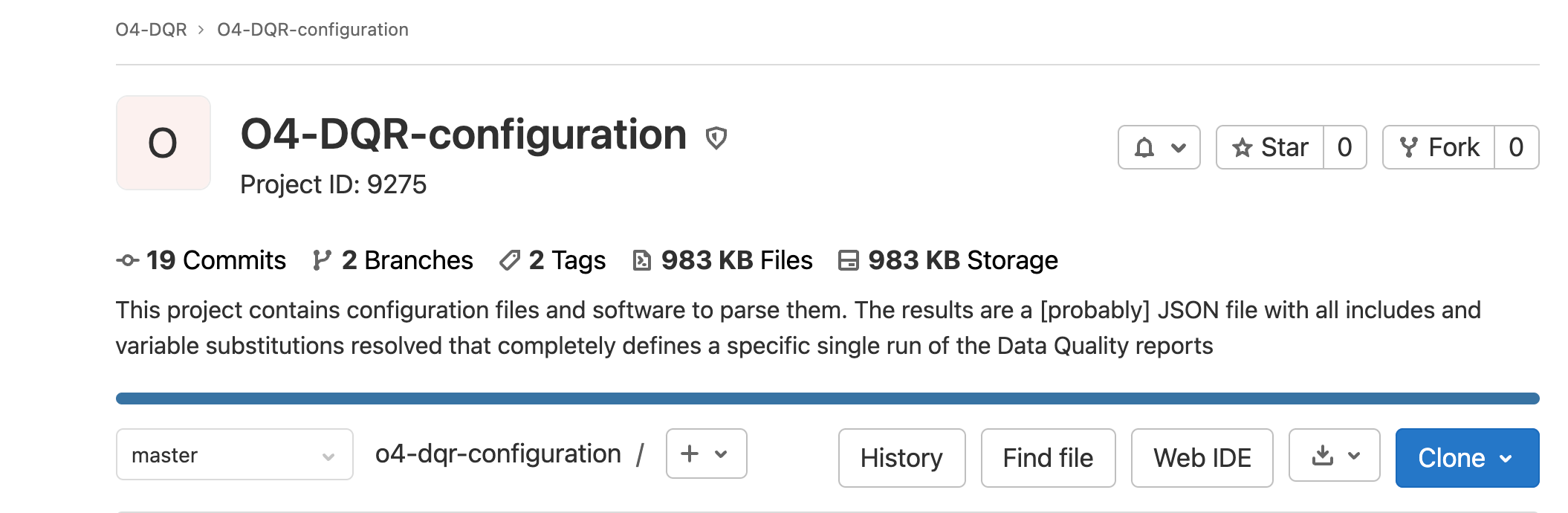
Fig. 2 Location of the fork and clone buttons.
# clone your fork to your local machine
git clone git@git.ligo.org:USERNAME/o4-dqr-configuration.git
Then to keep the project up to date set the upstream repo:
cd o4-dqr-configuration
# Add 'upstream' repo to list of remotes
git remote add upstream git@git.ligo.org:o4-dqr/o4-dqr-configuration.git
# Verify the new remote named 'upstream'
git remote -v
This produces:
$ git remote -v
origin git@git.ligo.org:joseph-areeda/o4-dqr-configuration.git (fetch)
origin git@git.ligo.org:joseph-areeda/o4-dqr-configuration.git (push)
upstream git@git.ligo.org:o4-dqr/o4-dqr-configuration.git (fetch)
upstream git@git.ligo.org:o4-dqr/o4-dqr-configuration.git (push)
The standard process keeps the master branch in the fork in sync with the upstream repo. Changes are made on a branch of the master branch. Ensure the master is up to date and create a new branch with the following set of commands:
git fetch upstream
git checkout master
git rebase upstream/master
git checkout -b my-new-branch
Add your new code to new branch. When you are ready to create a Merge Request:
# Update master branch.
git fetch upstream
git checkout master
git rebase upstream/master
# Rebase your branch to master
git checkout my-new-branch
git rebase master
# resolve any conflicts, hopefully none
# add any new files
git add <new file1> <new file 2>
# commit your changes
git commit --all
# the first time you push to your fork
git push --set-upstream origin my-new-branch
# subsequent pushes
git push
The above will start the CI/CD pipelines. Log into your project, choose the new branch and confirms the checks succeeded.
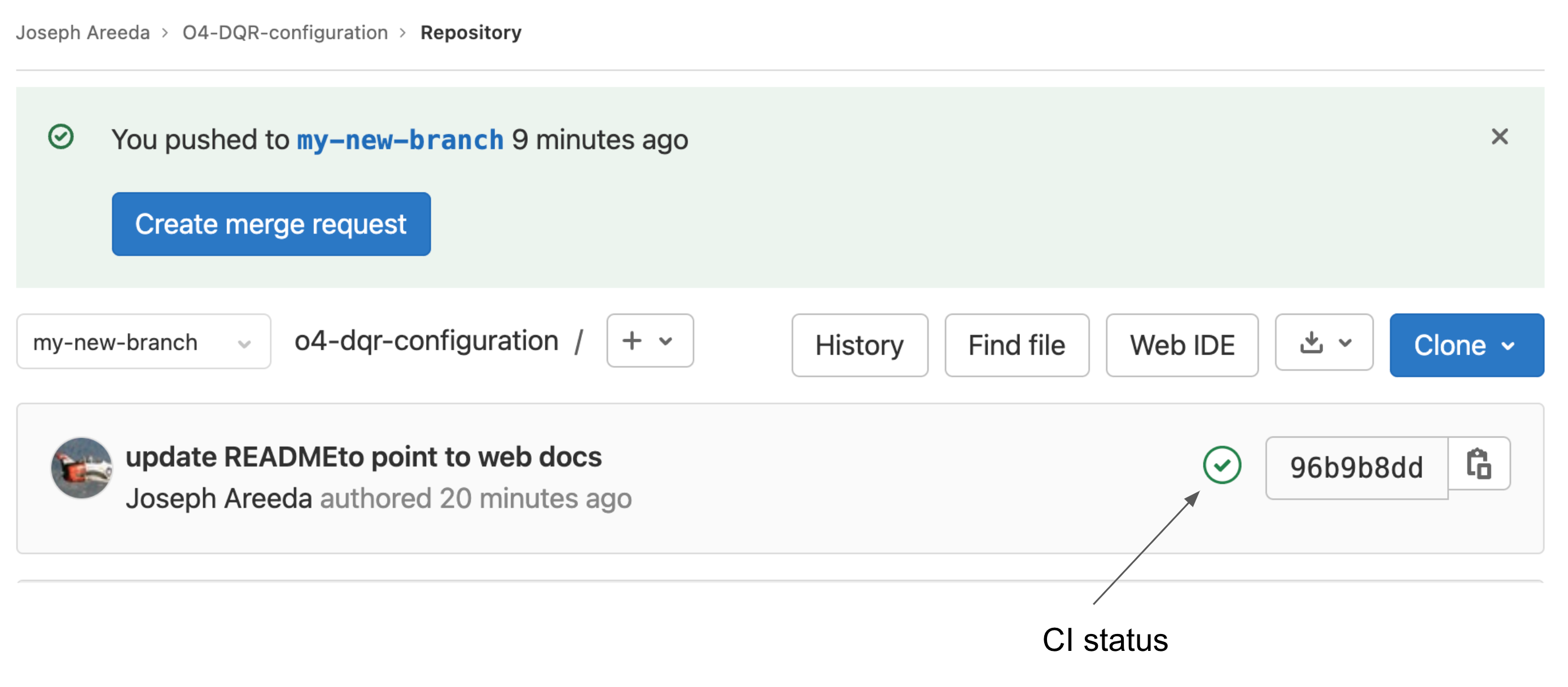
Fig. 3 Continuous Integration status and create merge request
The CI status indicator is either a green check mark or a red X. It is also a link to the pipline results.
Confirm the pipeline has passed, or identify a reason to update the pipeline.
Note
When changes are committed to a branch that is involved in an active Merge Request, the CI/CD pipelines are rerun. No further action is needed to update the MR.
Create the Merge Request. Please be clear why the changes were made. If the changes resolve one or more issues please include references. You may leave the Assignees, Reviewers and Milestone blank.
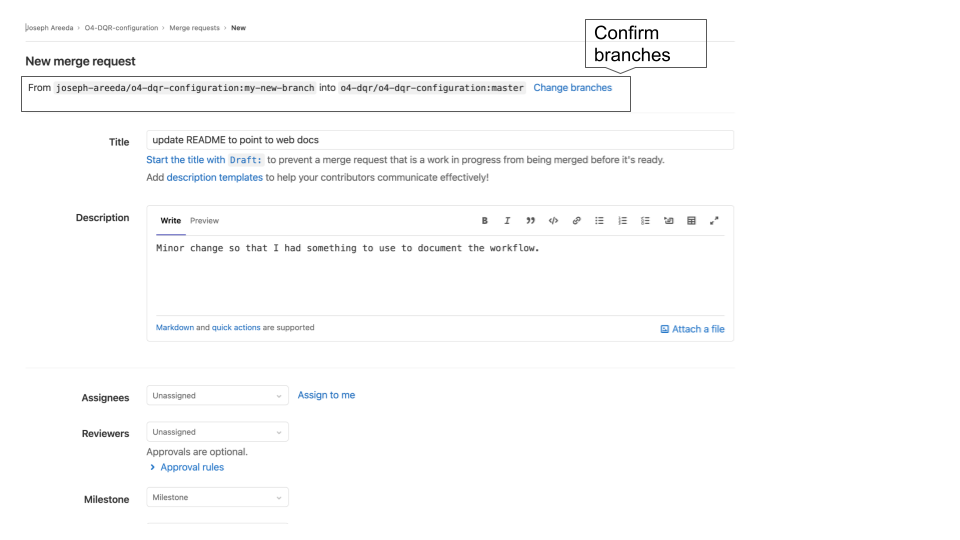
Fig. 4 Merge request dialog
Merge requests will be reviewed and approved by someone besides the author.
Delete branch after merge
It is customary, but not necessary to use a branch for a single task. Once a branch has been merged upstream or abandoned the branch can be deleted.
To delete a remote branch:
git push --delete origin my-new-branch
To delete the local branch:
git branch -d my-new-branch
git branch -D my-new-branch
-d is an alias for –delete and -D an alias for –delete –force
Note
Even deleted branches are kept in the git repository. If needed they can be recovered. See https://stackoverflow.com/questions/3640764/can-i-recover-a-branch-after-its-deletion-in-git